Technological progress
The rapid advances in cutting-edge technologies and informatics tools utilized in biomedical science to generate and process high-throughput biological datasets have finally spread into various economic sectors. Omic technologies adopt a holistic view of the molecular characteristics and actions that make up a cell, tissue or organism. “Omics” are novel comprehensive approaches for the analysis of complete genetic or molecular profiles of organisms by using integrative pipelines, showing how complex interactions between genes and molecules influence the phenotype, e.g. the disease symptoms in a patient. Great improvements in next-generation sequencing, microarray, mass spectrometry and nuclear magnetic resonance technologies have transformed biological and biomedical research over the past years. The analysis of omic data is a rapidly expanding field, with the constant development of new statistical methods and the creation of new perspectives not only for medical research and diagnosis. [1,2,3,4,5]
Examples of different types of omics: [2,3,4,5]
- Genomics: entire set of genes, “the genetic landscape” of an organism
- Proteomics: entire set of proteins produced by an organism
- Transcriptomics: entire set of RNA molecules
- Pharmacogenomics: the effect of variations within the human genome on drug response
Compared to single omics interrogations, multi-omics can provide researchers with a greater understanding of the flow of information, from the original cause of disease (genetic, environmental or developmental) to the functional consequences or relevant interactions. [2,3]
Computational Biology
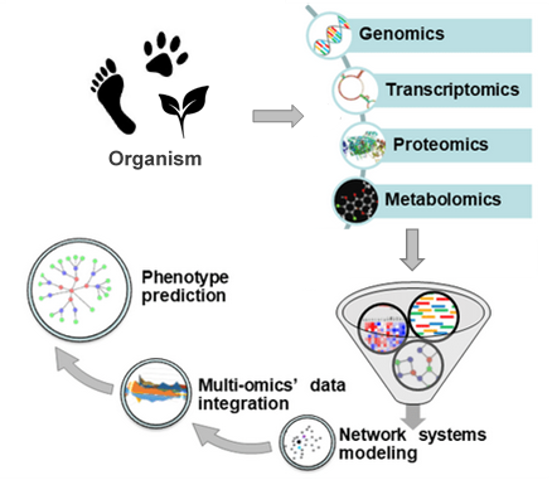
Bioinformatics uses advanced computational tools for the management and analysis of biological data that are mined from large databases. The combination of bioinformatics tools with adequate databases allows the generation of maps of cellular and physiological pathways. This integrative approach is called computational biology. Bioinformatics techniques enable a complete representation of the cell and the organism as well as the prediction of highly complex systems like cellular interaction networks or the phenotypes of organisms (Fig. 1). [6]
Merging omics and bioinformatics provides the basis of systems biology, a study field used in modeling organisms to enhance the understanding of the complex biological interactions occurring within cells and tissues at the gene, protein and metabolite level. [2,5] But with decreasing time and cost effort to generate these datasets, omics data integration has created both exciting opportunities and immense challenges for biologists, computational biologists, biostatisticians and biomathematicians.
Challenges
Nowadays, there are plenty of web-based solutions for data storage, sharing and analysis. Examples of common portals to access and download omic data are Gene Expression Omnibus, ArrayExpress, Expression Atlas or Ensembl. Other sites provide a framework for analyzing omics data in an interactive and multi-layered fashion, such as NCBI, UCSC or Human Brain Atlas. The rise of a high number of bioinformatics tools has fostered initiatives aimed at generating portals to list them and support their effective use. For example, EBI has a bioinformatics service portal listing a variety of databases and tools tailored for specific quests or topics, while OMICtools is a library of software, databases and platforms for big-data processing and analysis. [4,5]
But although there are various systems biology tools and applications available for network analysis, pathway construction, genome alignments, visualization and many more tasks, hardly any of these approaches can integrate three or more omics datasets. Managing and integrating such multi-dimensional data continues to be difficult, partly because each omics analysis can generate tera- to peta-byte sized data files daily. Thus, current biological and technical challenges include differences in data cleaning, storage and processing, biological contextualization, statistical validation, computational power and capacity as well as the lack of robust pipelines to integrate additional types of data. Innovative techniques of omic data integration are urgently needed for a broad range of research areas like food and nutrition science, analysis of microbiomes, genotype–phenotype interactions, systems biology and disease biology. [3,4,5]
Importance of omic tools in medicine
Comprehensive profiling has provided deep insight into the origin of diseases like cancer, coronary or infectious diseases, the search for diagnostic markers, potential therapies and the prediction of treatment response. Predictive markers or early diagnostic markers are of great benefit especially in cancer therapy, as various cancer types are detected at an advanced stage and 5‐year survival is poor. [1,5]
Multi-omic integrative analyses are meant to provide a comprehensive view of disease mechanisms that disrupt normal cellular functions and lead to disease progression and drug resistance. While computational and statistical analyses of single-omic datasets are well established, approaches for integrating multi-omics data are still far from being standardized. To keep up with the pace of data generation and growth of biological knowledge, existing methods should be extended or generalized, and new computational tools need to cope with the complexity and multi-level structure of the available information. [7]
Importance of omic tools in other sectors
Not only the medical sector benefits from specialized and integrated bioinformatics tools, but also other important fields like agriculture, plant science and animal science. As an example, honey bees are essential pollinators and crucial to the world’s agriculture. The health of honey bees has been declining over the past decade, with beekeepers losing more than a quarter of their colonies each winter since 2007. The causes of bee declines are complex, variable over space and time, and often difficult to identify.
The Canadian project BeeCSI, which was started in 2018, uses genomic tools to develop a new health assessment and diagnosis platform powered by stressor-specific markers. The project leaders working at York University and University of British Columbia join a cross-nation team of researchers, informaticians, beekeepers and diagnostic labs to develop a new tool which will be used to assess and diagnose honey bee health. The aim of BeeCSI is to bring industrial modernization in the form of this innovative tool which will give a quick diagnosis of bee health within living colonies, thus allowing beekeepers to immediately take appropriate measures to reduce honey bee loss. The 10 Mio $ project is funded by Genome Canada and Ontario Genomics. [8]
Future aspects
Studies in genomics, transcriptomics and proteomics have shaped our understanding of cellular complexity and heterogeneity. In the case of medicine, as the costs of omic analyses continually decrease, more types of high throughput data can be integrated into the clinic for individualized treatment regimens. Multi-scale omic data generation, development of analytical methodology, adaptation of those methods to specific disease, repeating this process for multiple diseases and integrating between them are the fundamental tasks of omic research, that can’t be conducted by only one research group. To ensure the proactive flow of data across and between different fields of expertise, these undertakings necessitate coordinated efforts of many skilled groups, leading to a standardization of data formats and pipelines for integrated multi-omic analyses. [1,2,4,5]
Contact person:




